|
|
def | __init__ (self, bnet) |
| |
| def | marginal (self, query) |
| | 0 = no change, 1 = update, 2 = retract isChange = 0 changedNodes = [] for (node, value) in zip(nodes, values): Make sure node has actually changed if not self.evidence[node.index] == value: changedNodes += node Check if node is retracted if not self.evidence[node.index] == -1: isChange = 2 break else: isChange = 1 More...
|
| |
|
def | global_propagation (self) |
| |
|
def | collect_evidence (self, prevClique, currentClique, sepset, isStart) |
| |
|
def | distribute_evidence (self, clique) |
| |
|
def | pass_message (self, fromClique, toClique, sepset) |
| |
| def | project (self, clique, sepset) |
| |
| def | absorb (self, clique, sepset, oldPotential) |
| |
|
def | build_join_tree (self, triangulatedGraph) |
| |
| def | create_sepset_priority_queue (self, cliques) |
| |
|
def | __init__ (self, bnet) |
| |
|
def | marginal (self) |
| |
This implementation of the Junction Tree inference algorithm comes from "Belief Networks: A Procedural Guide" By Cecil Huang an Adnan Darwiche (1996). See also Kevin Murhpy's PhD Dissertation. Roughly this algorithm decomposes the given bayes net to a moral graph, triangulates the moral graph, and collects it into cliques and joins the cliques into a join tree. The marginal is then computed from the constructed join tree.
| def PBNT.Inference.JunctionTreeEngine.marginal |
( |
|
self, |
|
|
|
query |
|
) |
| |
0 = no change, 1 = update, 2 = retract isChange = 0 changedNodes = [] for (node, value) in zip(nodes, values): Make sure node has actually changed if not self.evidence[node.index] == value: changedNodes += node Check if node is retracted if not self.evidence[node.index] == -1: isChange = 2 break else: isChange = 1
Just to avoid import errors assert(1 == 1) Do a global update for node in changedNodes: Just to avoid import errors assert(1 == 1) Update potential X and its likelihood with the new observation Then do global propagation (if only 1 cluster affected only have to distribute evidence. elif isChange == 2: Do a global retraction: Encode the new likelihoods (and do observation entry), Reinitialize the join tree, do a Global propagation. Just to avoid import errors assert(1 == 1)


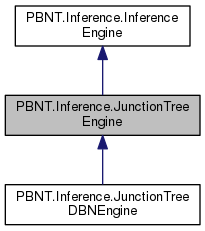
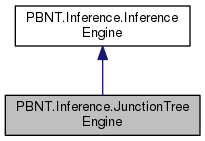
 Public Member Functions inherited from PBNT.Inference.InferenceEngine
Public Member Functions inherited from PBNT.Inference.InferenceEngine Public Attributes inherited from PBNT.Inference.InferenceEngine
Public Attributes inherited from PBNT.Inference.InferenceEngine 1.8.11
1.8.11